PAUNOV RESEARCH GROUP regularly has paid and unpaid vacancies for research assistants фор for both NU and non-NU students. Please contact Prof Paunov for vacancy information.
Please check on the Current Projects page the active projects which you may be interested to join as a research assistant.
OPPORTUNITIES FOR SELF-FUNDED PHD STUDENTS
At present we offer only the following PhD projects for self-funded post-graduate students. If you have your own funding or externally sponsored PhD studentship and are interested in some of the projects presented below, please contact Prof Paunov directly:
Project 1: Release kinetics of low soluble drugs encapsulated in sporopollenin microcapsules
Project 2: Whole cell biosensors for testing toxicity of nanomaterials and environmental contamination
Project 3: Fabrication of pH-responsive bacterial cellulose gels
Project 4: Development of novel nanoantibiotics based on a nanoimprinting of microbial cells
Project 1: Release Kinetics of Low Soluble Drugs Encapsulated in Sporopollenin Microcapsules
(for self-funded PhD students)
Supervisor: Prof Vesselin N. Paunov, Department of Chemistry
The outer layer (exine) of the pollen grain of mosses and ferns is composed largely of a substance known as sporopollenin1 which has been described as “one of the most extraordinary resistant materials known in the organic world”. Sporopollenins are probably the most resistant organic materials of direct biological origin found in nature and in geological samples” as intact spores are found in ancient sedimentary rocks, which are at least 500 million years old. In this project we will use a simple and robust technique for loading sporopollenin of Lycopodium Clavatum with inorganic or organic nanoparticles synthesised in situ. The idea of this method is to use the sporopollenin capsules as chemical micro-reactors where a chemical reaction is used to generate the product (e.g. of low solubility) inside the sporopollenin shells. Our approach will allow the sporopollenin capsules to be loaded with inorganic or organic materials produced as a result of a chemical reaction conducted selectively in the capsule interior. Using this method we will produce sporopollenin capsules filled with low-soluble forms of ibuprofen and other drugs of common interest, like the Parkinson drug Levodopa. We will study the rate of release of ibuprofen and levodopa from the sporopollenin capsules by using spectroscopic techniques. These have the potential for oral delivery of food supplements and drugs that are usually delivered by injection, more exactly target certain drugs and/or protect an active ingredient so that it can be delivered to the lower gut. The oral delivery is based upon the fact that certain plant pollens or other spores are capable of crossing the gut wall, largely intact, but is then destroyed within the blood stream, thereby releasing its contents.
Within this PhD project we will also be developing methods for triggered release of medicines encapsulated in these bioinspired microcapsules which rely on and external stimulus, like the change of pH, salinity and temperature for the release of the encapsulated material into the surrounding media.

The work will be done in the Dept. of Chemistry and will be supervised by Prof. Vessselin Paunov. For further details please contact Prof Paunov directly.
Project 2: Whole cell biosensors for testing toxicity of nanomaterials and environmental contamination
(for self-funded PhD students)
Supervisor: Prof. Vesselin N. Paunov, Department of Chemistry
At present, there are over 7 million chemical compounds reported in the specialised literature many of which are widely used on industrial scale in many consumer products and formulations. Only around 7000 compounds have ever been tested for genotoxicity and cytotoxicity. On the other hand, nanomaterials such as nanoparticles, nanodots, nanotubes and nanocomposites are also rapidly finding their way into many industrial or consumer products. However, the present nanomaterials, especially the ones made of metals, semiconductors and various inorganic compounds, pose significant environmental hazard post-use as they persist in the environment and over time may accumulate in humans and affect numerous live organisms. Testing of toxicity effects of chemicals and nanoparticle formulations can be a long and expensive process and usually involves controversial practices like using experiments on animals.
The present PhD project is devoted to the development of cheap, rapid and reliable methods for testing of industrial formulations and nanomaterials for potentially hazardous effects which is based on using whole bacterial cells as reporters for toxicity. Bacterial cells can be genetically reprogrammed to undertake novel tasks such as sensing specific chemicals, producing proteins and drugs. For example, a DNA-repair-competent strain of the brewer’s yeast S. Cerevisiae modified with a Green Fluorescent Protein (GFP) promoter can be used to test the genotoxicity effect of various chemicals which may interfere with the bacterial cell DNA. The bacterial DNA-repair gene gets activated upon mutagenic changes and this also unlocks the synthesis of GFP which presence can be detected by fluorescence microscopy. In analytical terms, the yeast cells start fluorescing soon after they get exposed to potentially mutagenic agents. Similar effect is expected when such cells are incubated with nanoparticles, some of which may have the ability to penetrate through the cellular membrane and affect the cell DNA.
We plan to functionalise GFP-reporter yeast cells with magnetic nanoparticles which would allow us to hold and position them into microfluidic devices (like the one bellow) where they will be rapidly incubated into solutions of the tested compounds and nanoparticles at different concentrations. The magnetic yeast cells will be hold onto the chip by a magnetic field.

The fluorescence intensity of the bacterial cells will be proportional to the activity of the gene responsible for DNA repair which would allow us to determine the concentration threshold of toxicity of the studied compound. Control experiments will be run with harmless materials for comparison purposes. After the test, the magnetic field is removed and the used cells will be replaced with a fresh sample of cells for further testing. The same test would also hold very well for examining the effect of nanoparticles on the cell viability and/or DNA-repair activity. It is expected that the use of whole bacterial cells as biosensors for toxicity of nanomaterials will enable us to achieve a rapid testing at minimal cost which would help us to develop a device for industrial standard in this area. We have some preliminary results obtained in our department which will be a good starting point for more focused effort of the PhD Student to do these studies for a variety of dispersed nanomaterials. Note that the proposed research uses bacterial cells for all tests and does not involve experiments with animals.
We also plan to use whole cell biosensors to assess potential contamination levels of soils and river sediments with potentially hazardous chemicals and nanomaterials. We will choose genetically modified reporter cells which have genes responding to a specific compound in their environment, coupled with a GFP promoter which starts expressing Green Fluorescent Protein when the gene processing specific compound is activated. Through collaboration with Kroto Institute at Sheffield University we have access to bacterial strains processing toluene/xylene, alkanes, salicilates and many others. We will magnetically functionalise such bacterial reporter cells and will release them into the soil or the sediment sample where they will be in contact with the materials and compounds embedded into their pores. After exposure, we will be collecting the magnetised reporter bacterial cells with a magnet and will be analysing the intensity of their fluorescence (or bioluminescence). The presence of the contamination of particular class of compounds will be detected from the level of GFP fluorescence. By using control experiments we will be able to calibrate the intensity of fluorescence (or bioluminescence) of the magnetic reporter cells to the exact level of contamination of the soil/sediment with the analyzed material.

Acinetobacter reporter cells decorated with magnetic nanoparticles allow their release and subsequent collection from the environment.
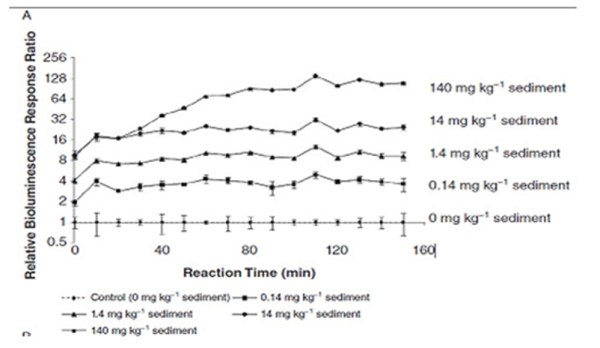
We have preliminary data obtained in collaboration of our group at the Department of Chemistry of the University of Hull and Kroto institute at Sheffield University which show that this technique works for salicilates as a model contaminant, as shown in the graph above where the cell bioluminescence was found proportional to the concentration of salicilates in the sediment sample. Within this PhD project we are also planning to do these tests for a variety of contaminants including nanoparticles of metals and semiconductors which would enable us to develop specific tests for assessment of contamination of soils, water tables and river sediments. These tests will then be performed in environmental conditions.
For further details please contact Prof Paunov directly.
Project 3: Fabrication of pH-Responsive Bacterial Cellulose Gels
(for self-funded PhD students)
Supervisor: Prof Vesselin N. Paunov, Department of Chemistry
Bacterial cellulose is a hydrogel formed by nano-fibres of cellulose in aqueous media which are a result of the continuous secretion of cellulose out of well known harmless bacteria, called Acetobacter Xylinum. The bacterial cellulose is non-toxic and fully biocompatible which gives it many practical applications, including food stuff (nata de coco), medical implants, etc. Since cellulose is a non-ionic polymer, pH changes in the aqueous phase have negligible effect on the swelling of the bacterial cellulose gels. On the other hand, microgels based on polyacrylic acid are known to increase their size in aqueous dispersion many times upon increase of pH due to the dissociation of the COOH-groups onto the microgel particle. The microgel swelling behaviour is also sensitive to the salt concentration as the salt ions screen out the electrostatic repulsion between the ionised COOH-groups of the microgel particles.
In this project we aim to integrate a special type of pH-sensitive microgel particles within the matrix of bacterial cellulose gels which would make them responsive to changes in the surrounding pH. The project will allow us to produce novel biocompatible materials which respond to pH trigger and are suitable for controlled release of encapsulated active components.
We will study the degree of pH-induced swelling of bacterial cellulose gels after loading with pH sensitive microgels. Our aims will be to find appropriate microgel and salt concentration range for optimal loading into the bacterial cellulose gel and to study the rate of release of fluorescent marker molecules encapsulated in the microgel particles. We will monitor both the pH and the concentration of released fluorescent tracer in order to understand the release kinetics of the microgel entrapped in the bacterial cellulose matrix.
(a)  (b)
(b) 
(a) Nanofiber network of bacterial cellulose. Electron micrograph, magnification 1000. (b) Typical bacterial cellulose gel (nata de coco).
The work will be done in the Department of Chemistry and will be supervised by Prof. Paunov.
For further details please contact Prof Paunov directly.
Project 4: Development of Novel Nanoantibiotics Based on a Nanoimprinting of Microbial Cells
(for self-funded PhD students)
Supervisor: Prof Vesselin N. Paunov, Department of Chemistry
This proposal is devoted to the development of novel class of nanoparticles capable of deactivating specific pathogenic organisms like viruses and bacteria. We call this novel class of agents “nanoantibiotics” and the new idea here is to use nanoimprinting technology to produce a “negative” inorganic replica of the pathogen that fits specifically to its shape and functionality. For example, once in contact with the virus pathogen, the nanoantibiotic particle can bind specifically to it and can prevent the viral capsid (from disassembling and releasing the viral DNA). Similarly, nanoantibiotics designed for bacterial pathogens can “fit” on part of the bacterial wall and stop its further mitosis and reproduction. Over the last decade nanoimprinting techniques have been successfully used for development of molecular sensors and miniaturisation of electronic circuits. Recently, there have been several fundamental developments involving the nanotemplating and assembly of nanoparticles in liquid surfaces. It has been demonstrated that nanoparticles can be superior emulsion stabilisers in the absence of any surfactants thanks to their adsorption at the emulsion interface drops.[1] Noble, Paunov, et al.[2] developed further this idea by templating water-in-oil emulsions stabilised with nanoparticles to produce colloidosomes (hollow shell capsules with a membrane consisting of a particle monolayer). Recently Paunov et al developed the Gel Trapping Technique (GTT) [3,4] which proved to be a very powerful method not only for determining contact angles of individual microparticles at liquid surfaces but also for synthesis of novel materials, like asymmetrically functionalised microparticles. This technique involves spreading the particles at air-water or oil-water interface and subsequent gelling of the aqueous sub-phase with a non-adsorbing hydrocolloid. This allows the adsorbed particles to be “trapped” at the liquid surface and the top phase (oil or air) to be removed and replaced with curable silicone (PDMS). After curing, the solid PDMS replica is peeled off the gel and the exposed part of the particle monolayer can be modified to produce asymmetric functionality. For example, exposure to gold vapour over polystyrene latex particle monolayer “trapped” on PDMS allows the deposition of gold film over the exposed part of the particle surface, which can be further released to produce “capped” particles by stretching the PDMS resin. Further, the latex particles can be dissolved to produce golden “cap” particles that can match and fit on their original templates when mixed together with the latex suspension.

Figure 1. Schematics of the nanoimprinting method for preparation of nanoantibiotics based on viral monolayers picked up from air-water interface by using the Gel Trapping Technique. The nanoimprints of the viruses are produced by breaking down of gold sputtered film by stretching of the resin and ultrasonic treatment.
To produce nanoantibiotics and demonstrate their functionality we will follow two main avenues:
(i) Gel trapping method: Here we would like to explore further this idea to create novel class of antibiotic agents based on asymmetric nanotemplating of the respective pathogens (viruses or bacteria) by using the GTT and producing nanoparticles that are “tailor made” to cap them up to achieve deactivation. We are planning to produce monolayers of viruses at air-water or oil-water interface. This can be done under controlled conditions in a Langmuir through at constant surface pressure or alternatively by using a spreading solvent in a Petri dish with aqueous solution. We are planning to test the method with several types of viruses of different shape and functionality, e.g. tobacco mosaic virus. Very similar scheme will be applied to monolayers of bacteria at a liquid surface. The viral monolayer will be templated with GTT (see ref 4) and will be nanoimprinted with gold vapour within the monolayer. The gold nanocoating which imprints the exposed part of the viruses will be broken down to small nanoparticles by stretching of the PDMS resin and use of ultrasound treatment in surfactant solution followed by bleaching of the organic residues from the viruses in weak solution of hydrogen peroxide. The nanoshells can be further functionalised by the thiol reaction and grafting of specific polysaccharides for higher specificity of binding to the target viruses. The efficiency of the produced nanoantibiotics will be tested by mixing with small amounts of virus sample, which is expected to result in specific irreversible binding (capping) and deactivation of the viruses by the nanoantibiotic particles (see Figure 1).

Figure 2. Schematics of the Broken Shell Method for preparation of nanoantibiotics. The method is illustrated for viral particles but can also be applied to bacteria. The viruses are used as templates for deposition of inorganic shell, e.g. CaCO3 followed by ultrasonic treatment to break the produced nanoshells. The removal of the viral templates and grafting of surface recognition groups produces nanoparticles capable of recognising both the shape and the surface chemistry of the virus and can deactivate it by binding to it irreversibly and preventing it from entering the cell. In the case of bacteria, the nanoantibiotics would stop the bacteria from further mitosis.
(ii) Broken Shell Method. In this avenue of research we would like to explore the possibility of nanoimprinting of viruses and bacterial pathogens by nanoimprinting them in the bulk with a nanoshells of inorganic material (calcium carbonate or silica). These materials are well known to replicate extremely well the morphology and functionality of the surface of the templates (e.g. viruses) down to molecular level. The templated particles will be broken to pieces by application of ultrasound and the organic materials (the templates) will be bleached by using hydrogen peroxide or other oxidising agent. Following a similar scheme as in the Gel Trapping Method (see above) we will further improve the functionality of the obtained nanoshells by grafting specific polysaccharides to their surface, so that they will be able to “fit and bind” perfectly to the target pathogenic particles (viruses and bacteria). Control experiments will be done with a series of different viruses and bacteria (and mixtures of them) to show the proof of principle for the antibiotic action of these nanoshells (see Figure 2).
Such “tailor made” nanoshells are expected to act as powerful antibiotics at very low concentration since one nanoantibiotics particle would be able to deactivate a whole virus or specific bacteria. The lack of other organic materials in the composition of the nanoantibiotics particles and the low concentrations needed for their antibiotic action are expected to lead to low toxicity and high efficiency against the specific pathogens they are designed to deactivate. The use of calcium carbonate (natural material) for the nanoshells would also lead to a lack of side effects. It is anticipated that this novel class of antibiotic nanoparticles that are “tailor made” for specific pathogens can be applied to highly resistant lines of bacteria (like the super bug MRSA) and many others where most conventional antibiotics and powerless. The improvement here is that the nanoantibiotics particles would recognise not only the surface chemistry of the pathogen but also its shape as a result of the nanoimprinting process. It is expected that these nanoantibiotics can also be used as non-toxic bacteriostatic agents, which would make them applicable in the food industry, for example to prevent E-coli bacteria from developing and propagating on food formulations.
- Borovicka, J., Stoyanov, S.D., Paunov, V.N., Shape recognition of microbial cells by colloidal cell imprints, Nanoscale, 57 (2013) 8560 – 8568.
- Borovicka, J., Metheringham, J., Madden, L.A., Walton C.D., Stoyanov, S.D., Paunov, V.N., “Photothermal colloid antibodies for shape-selective recognition and killing of microorganisms.”, J. Am. Chem. Soc., 135 (2013), 5282-5285. [Highlighted by ACS Press]
For further details please contact Prof Paunov directly.
